|
2/17/2023 0 Comments Macvector plasmid mappingThe protein and its bound DNA sequences were separated from unbound oligonucleotide by EMSA. The oligonucleotides, which were end-labeled with 32P, were 55 bp in length and contained a central random stretch of 15 bp. To test the possibility that NK2-SD plays a role in DNA binding, we expressed in Escherichia coli a protein containing the homeodomain and NK2-SD of Nkx2.2 (amino acids 112–222) and used the purified protein to select for binding of specific DNA sequences from among a library of oligonucleotides. Indeed, the structure of the NK2-SD suggests a single loop reminiscent of the nonhelical loops in known DNA-binding domains ( 8). Previous studies identifying the optimal binding sites of NK-2 homeodomain proteins have used protein constructs containing the homeodomain alone ( 9), and none have included the NK2-SD. Experimental Procedures Binding Site Selection. Our results suggest a model in which Nkx2.2, regulated by interactions through the NK2-SD, causes activation of specific genes during development. The NK2-SD does contribute significantly, however, to transactivation function, inasmuch as mutation or deletion of this domain unmasks a powerful transcriptional activation function in Nkx2.2. Our results reveal that Nkx2.2 recognizes the DNA sequence T C TAAGT G A G CTT and that the NK2-SD does not contribute significantly to the specificity of DNA binding. We determined the DNA-binding characteristics and transactivation properties of Nkx2.2 as they relate to the NK2-SD. However, the direct target genes and the specific function of Nkx2.2 are unknown. The targeted disruption of Nkx2.2 alters the development of the spinal cord ( 13) and pancreatic islet cells ( 14). Nkx2.2 was originally identified as a member of the NK-2 class of transcription factors expressed in the developing central nervous system ( 11) and pancreas ( 12). In this study, we used Nkx2.2 as a model protein for investigating the functional role of the NK2-SD. However, these studies did not implicate the TN domain and NK2-SD directly in the function of these proteins ( 9, 10). Previous studies have mapped the functional domains of Nkx2.1 and Nkx2.5 and have found both activation and repression domains in both proteins. This structure suggests that the NK2-SD might function as an accessory DNA-binding domain or as a protein–protein interaction interface ( 8). The NK2-SD is composed of a hydrophobic core sequence VAVPVLV, with a central proline that prevents helix formation, and is flanked by basic amino acids. The NK2-SD is unique to NK-2-class proteins and is separated from the C-terminal end of the homeodomain by a short linker. The short TN domain is found at the N-terminal region of most NK-2 proteins. The homeodomain is a highly conserved DNA-binding domain the homeodomains of the NK-2 class are believed to recognize the DNA sequence 5′-CAAG-3′, whereas most other homeodomain transcription factors recognize 5′-TAAT-3′ ( 6, 7). The transcription factors of this class contain three highly conserved regions: the homeodomain, the TN (tin) domain (or NK decapeptide), and the NK2-specific domain (NK2-SD also called the NK2 domain) ( 1). These results demonstrate that the NK2-SD functions as an intramolecular regulator of the C-terminal activation domain in Nkx2.2 and support a model in which interactions through the NK2-SD regulate the ability of NK-2-class proteins to activate specific genes during development. The NK2-SD also can mask transactivation from the paired homeodomain transcription factor Pa圆, but it has no effect on transcription by itself. Interestingly, this C-terminal region functions as a transcriptional activator only in the absence of an intact NK2-SD. 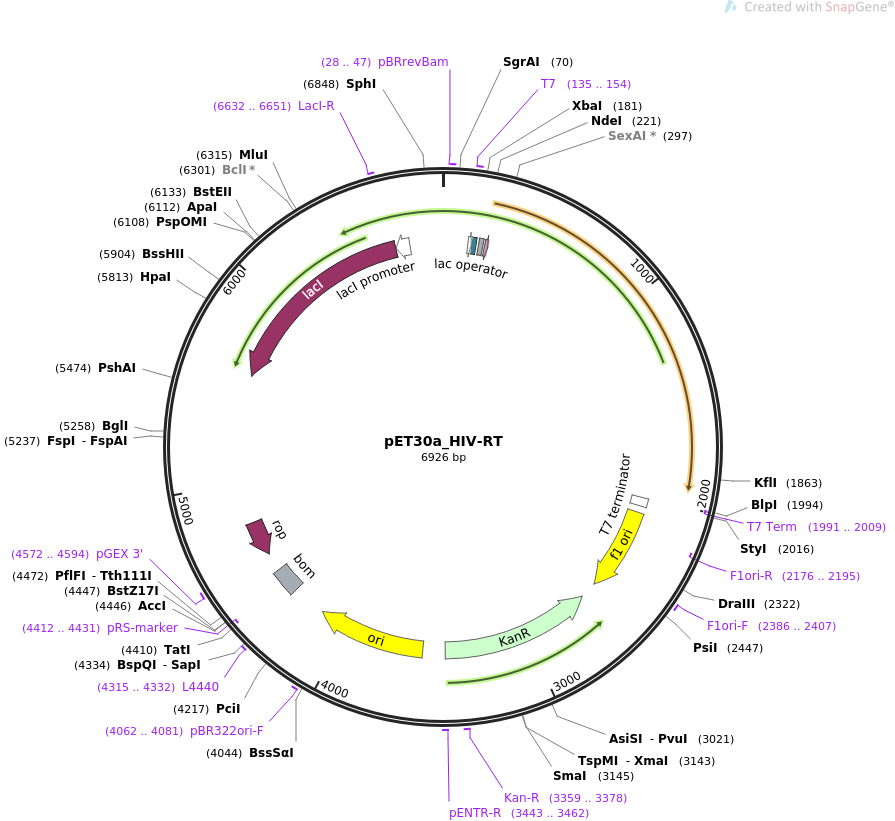
To determine whether the NK2-SD contributes to transactivation, we used GAL4-Nkx2.2 fusion constructs to map a powerful transcriptional activation domain in the C-terminal region beyond the conserved NK2-SD. The consensus sequence (T C TAAGT G A G CTT) is similar to the known binding sequences for other NK-2 homeodomain proteins, but we show that the NK2-SD does not contribute significantly to specific DNA binding to this sequence. To assess the possibility that the NK2-SD may contribute to DNA-binding specificity, we used a PCR-based approach to identify a consensus DNA-binding sequences for Nkx2.2, an NK-2 family member involved in pancreas and central nervous system development. 
The primary structure of the NK2-SD suggests that it might function as an accessory DNA-binding domain or as a protein–protein interaction interface. The function of this domain, however, remains unknown. The developmentally important homeodomain transcription factors of the NK-2 class contain a highly conserved region, the NK2-specific domain (NK2-SD).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
